Nanoconfinement
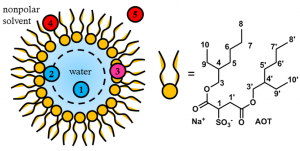
For more than the past two decades, the Levinger group has explored the impact of nanoconfinement on liquids. As a leader in the field, we have grappled with questions ranging from concepts of molecular motion in confinement to understanding what bulk properties like pH and ionic strength mean in these nanoscopic proportions.
We use reverse micelles as a model system to study nanoconfined liquids. At appropriate proportions, these nanodroplets form spontaneously in ternary mixtures of non-polar, polar, and amphiphilic molecules. Sometimes an additional material, a cosurfactant, is needed to stabilize the nanodroplet reverse micelles. Our work on these systems enlists ultrafast laser spectroscopy, solvation dynamics, time-resolved infrared spectroscopy, as well as NMR spectroscopy, including 1D, 2D and multinuclear methods.
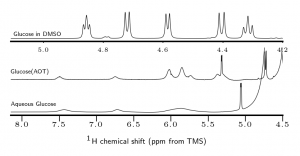
NMR spectra of glucose in DMSO, glucose in AOT reverse micelles, and glucose in aqueous solution. The reverse micelle environment slows chemical exchange with water making hydroxyl signals appear.
Our current studies focus on the role and impact of carbohydrates in nanoconfinement. We have demonstrated interesting behavior when glucose, sorbitol and other carbohydrates are encapsulated in reverse micelles. Specifically, we observe smaller reverse micelle sizes for samples prepared with aqueous solution compared to those prepared with water. Nanoconfinement also has a strong impact on chemical exchange between carbohydrate hydroxyl groups and water molecules, slowing the rate of exchange by orders of magnitude.
We continue to explore how carbohydrates impact reverse micelles and how the nanoconfinement in reverse micelles affects the carbohydrate properties. Questions we seek to addresss include:
- How does a confining environment affect molecular motion?
- How do carbohydrates impact water in a confined environment?
- Most of our studies have focused on glucose. Are results transferable to other carbohydrates?
- How does concentration affect exchange dynamics?
- How does surfactant affect exchange dynamics?
We use ultrafast laser spectroscopy to measure solvation dynamics in reverse micelles.
Cryopreservation
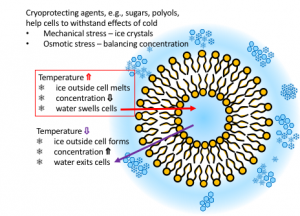
A new direction in the Levinger group addresses cryopreservation. How do cells stay alive under extreme conditions? Many organisms have developed mechanisms that protect them from cold and dry environments. These organisms display similar motifs, such as antifreeze proteins, that allow them to withstand low temperatures. Scientists have found artificial methods that work to preserve some biological materials, e.g., cells and tissue, at low and cryogenic temperatures.
Artificial cryopreservation methods are critical in a wide range of applications from fertility clinics, animal breeding, to preservation of plant material for agriculture. Cryopreservation formulations that are effective for preserving one specific cell type often perform worse or not at all when applied to different cell types, cells from different species, and sometimes even between individual animals of the same genus and species.
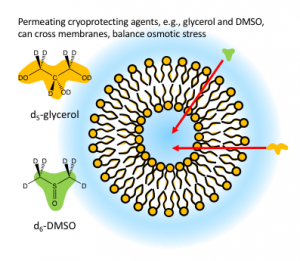
We are developing a program to explore fundamental molecular and cell properties that lead to successful cryopreservation.The empirical approach to artificial cryopreservation that has led to formulations currently in use means that although we know what can work to preserve some samples, we do not have foundation understanding about why or how these formulations work. Without this understanding, it is hard to predict alternative formulations for different organisms.
Our studies use stimulated Raman microscopy to measure properties of permeating cryopreservation agents, such as glycerol or DMSO, as they interact with cells and model vesicle systems. We measure properties such as how fast cryoprotectants cross into cells, where cryoprotectants reside in cells, cooperativity between cryopreservation agents. Through these studies, we aim to develop foundational knowledge that will enable us to predict effective cryopreservation formulation for a wide range of systems including plants and animal cells.
